Abstract
This protocol describes the process of isolating and engineering antibodies or proteins for increased affinity and stability using yeast surface display. Single-chain antibody fragments (scFvs) are first isolated from an existing nonimmune human library displayed on the yeast surface using magnetic-activated cell sorting selection followed by selection using flow cytometry. This enriched population is then mutagenized, and successive rounds of random mutagenesis and flow cytometry selection are done to attain desired scFv properties through directed evolution. Labeling strategies for weakly binding scFvs are also described, as well as procedures for characterizing and 'titrating' scFv clones displayed on yeast. The ultimate result of following this protocol is a panel of scFvs with increased stability and affinity for an antigen of interest.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
06 September 2007
In the version of this article initially published, the x-axis of Figure 4b should have been labeled “PE antigen binding”, not “Alexa Fluor 488 (c-Myc)”. The figure has been corrected in the HTML and PDF versions of the article.
References
Kieke, M.C., Cho, B.K., Boder, E.T., Kranz, D.M. & Wittrup, K.D. Isolation of anti-T cell receptor scFv mutants by yeast surface display. Protein Eng. 10, 1303–1310 (1997).
Colby, D.W. et al. Development of a human light chain variable domain (VL) intracellular antibody specific for the amino terminus of huntingtin via yeast surface display. J. Mol. Biol. 342, 901–912 (2004).
Graff, C.P., Chester, K., Begent, R. & Wittrup, K.D. Directed evolution of an anti-carcinoembryonic antigen scFv with a 4-day monovalent dissociation half-time at 37 °C. Protein Eng. Des. Sel. 17, 293–304 (2004).
Razai, A. et al. Molecular evolution of antibody affinity for sensitive detection of botulinum neurotoxin type A. J. Mol. Biol. 351, 158–169 (2005).
Boder, E.T., Midelfort, K.S. & Wittrup, K.D. Directed evolution of antibody fragments with monovalent femtomolar antigen-binding affinity. Proc. Natl. Acad. Sci. USA 97, 10701–10705 (2000).
Holler, P.D. et al. In vitro evolution of a T cell receptor with high affinity for peptide/MHC. Proc. Natl. Acad. Sci. USA 97, 5387–5392 (2000).
Rao, B.M., Driver, I., Lauffenburger, D.A. & Wittrup, K.D. High-affinity CD25-binding IL-2 mutants potently stimulate persistent T cell growth. Biochemistry 44, 10696–10701 (2005).
Kim, Y.S., Bhandari, R., Cochran, J.R., Kuriyan, J. & Wittrup, K.D. Directed evolution of the epidermal growth factor receptor extracellular domain for expression in yeast. Proteins 62, 1026–1035 (2006).
Piatesi, A. et al. Directed evolution for improved secretion of cancer-testis antigen NY-ESO-1 from yeast. Protein Expr. Purif. (in the press).
Boder, E.T. & Wittrup, K.D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
VanAntwerp, J.J. & Wittrup, K.D. Fine affinity discrimination by yeast surface display and flow cytometry. Biotechnol. Prog. 16, 31–37 (2000).
Shusta, E.V., Kieke, M.C., Parke, E., Kranz, D.M. & Wittrup, K.D. Yeast polypeptide fusion surface display levels predict thermal stability and soluble secretion efficiency. J. Mol. Biol. 292, 949–956 (1999).
Feldhaus, M.J. et al. Flow-cytometric isolation of human antibodies from a nonimmune Saccharomyces cerevisiae surface display library. Nat. Biotechnol. 21, 163–170 (2003).
Hoogenboom, H.R. Selecting and screening recombinant antibody libraries. Nat. Biotechnol. 23, 1105–1116 (2005).
van den Beucken, T. et al. Affinity maturation of Fab antibody fragments by fluorescent-activated cell sorting of yeast-displayed libraries. FEBS Lett. 546, 288–294 (2003).
Weaver-Feldhaus, J.M. et al. Yeast mating for combinatorial Fab library generation and surface display. FEBS Lett. 564, 24–34 (2004).
Wang, X.X. & Shusta, E.V. The use of scFv-displaying yeast in mammalian cell surface selections. J. Immunol. Methods 304, 30–42 (2005).
Richman, S.A. et al. Development of a novel strategy for engineering high-affinity proteins by yeast display. Protein Eng. Des. Sel. 19, 255–364 (2006).
Boder, E.T. & Wittrup, K.D. Yeast surface display for directed evolution of protein expression, affinity, and stability. Methods Enzymol. 328, 430–444 (2000).
Feldhaus, M. & Siegel, R. Flow cytometric screening of yeast surface display libraries. Methods Mol. Biol. 263, 311–332 (2004).
Colby, D.W. et al. Engineering antibody affinity by yeast surface display. Methods Enzymol. 388, 348–358 (2004).
Siegel, R.W., Coleman, J.R., Miller, K.D. & Feldhaus, M.J. High efficiency recovery and epitope-specific sorting of an scFv yeast display library. J. Immunol. Methods 286, 141–153 (2004).
Yeung, Y.A. & Wittrup, K.D. Quantitative screening of yeast surface-displayed polypeptide libraries by magnetic bead capture. Biotechnol. Prog. 18, 212–220 (2002).
Zaccolo, M., Williams, D.M., Brown, D.M. & Gherardi, E. An approach to random mutagenesis of DNA using mixtures of triphosphate derivatives of nucleoside analogues. J. Mol. Biol. 255, 589–603 (1996).
Stemmer, W.P. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 (1994).
Swers, J.S., Kellogg, B.A. & Wittrup, K.D. Shuffled antibody libraries created by in vivo homologous recombination and yeast surface display. Nucleic Acids Res. 32, e36 (2004).
Chowdhury, P.S. & Wu, H. Tailor-made antibody therapeutics. Methods 36, 11–24 (2005).
Meilhoc, E., Masson, J.M. & Teissie, J. High efficiency transformation of intact yeast cells by electric field pulses. Bio/Technology 8, 223–227 (1990).
Gietz, R.D. & Woods, R.A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 350, 87–96 (2002).
Boder, E.T. & Wittrup, K.D. Optimal screening of surface-displayed polypeptide libraries. Biotechnol. Prog. 14, 55–62 (1998).
Altman, J.D. et al. Phenotypic analysis of antigen-specific T lymphocytes. Science 274, 94–96 (1996).
Acknowledgements
This work was supported by CA96504, CA101830 and AI065824 from the National Institutes of Health, a David Koch Graduate Fellowship from the Massachusetts Institute of Technology Center for Cancer Research (G.C.), National Defense Science and Engineering Graduate Fellowships (G.C. and B.J.H.), a National Institute of General Medical Sciences Biotechnology Training Grant (S.L.S.) and a National Science Foundation Graduate Fellowship (S.M.L.). We thank the Massachusetts Institute of Technology Flow Cytometry Core Facility for their assistance, and E. Boder, J. Cochran, M. Feldhaus, M. Roguska and E. Shusta for detailed feedback on this protocol manuscript.
Author information
Authors and Affiliations
Contributions
G.C., Introduction, labeling and enrichment protocols, Anticipated Results, Figs. 1, 2 and 4, and assembly of manuscript; W.L.L., Box 1, Fig. 3 and Supplementary Fig. 1; B.J.H., characterization, mutagenesis and transformation protocols and Fig. 3; S.L.S., MACS protocol; S.M.L., labeling and titration protocols, Figs. 4 and 5 and Table 1; and all authors participated in discussions on optimal protocol parameters and the editing and revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
pCTCON2 plasmid map. (PDF 92 kb)
Supplementary Table 1
Preferred yeast culturing vessels. (PDF 104 kb)
Rights and permissions
About this article
Cite this article
Chao, G., Lau, W., Hackel, B. et al. Isolating and engineering human antibodies using yeast surface display. Nat Protoc 1, 755–768 (2006). https://doi.org/10.1038/nprot.2006.94
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.94
This article is cited by
-
A fine-tuned yeast surface-display/secretion platform enables the rapid discovery of neutralizing antibodies against Clostridioides difficile toxins
Microbial Cell Factories (2023)
-
Immunotherapy in hematologic malignancies: achievements, challenges and future prospects
Signal Transduction and Targeted Therapy (2023)
-
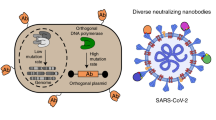
A universal reagent for detection of emerging diseases using bioengineered multifunctional yeast nanofragments
Nature Nanotechnology (2023)
-
High-throughput characterization of HLA-E-presented CD94/NKG2x ligands reveals peptides which modulate NK cell activation
Nature Communications (2023)
-
De novo design of modular peptide-binding proteins by superhelical matching
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.